
Kunio Miki started his career as a structural biologist 40 years ago as he was fascinated by the beauty of crystals, in which molecules are aligned in a regular pattern. In those days, it took an excessive amount of time for crystal structure analysis of proteins. It is hard to believe now but taking a couple of years to complete structure determination of one protein was just reasonable and not surprising at all.
Then, in 1982, he started working at Max Planck Institute of Biochemistry located near Munich, Germany, investigating the crystal structure of photosynthetic reaction center -- a key component for the photochemical conversion of light into chemical energy. Later in 1988, his co-workers were awarded the Nobel Prize in Chemistry as they were the first to succeed in elucidating the complete three-dimensional structure of a membrane protein complex by means of crystallography.
The 1990s witnessed a considerable advance in this field, thanks to the progress in genetic engineering, which enabled the preparation of sufficient amounts of protein samples for crystallization. The development of synchrotron radiation providing stronger X-ray source and broader spectrum also contributed to it.
The number of protein structures deposited in the Protein Data Bank (PDB) database has grown from only 500 in 1990 to close to 120 thousand today.
Life science in the 21st century started with structural genomics that seeks to describe the 3D structure of every protein encoded by a given genome. In Japan, this worldwide trend has led to a promotion of government-financed projects in this area, which helped speed up and generalize of the research method in protein crystallography.
It was on this background that the membrane-protein crystallography also made steady progress (Fig. 1) and it yielded an innovative field of ultrahigh resolution structure analysis and charge-density study in protein crystallography.
In such ultrahigh resolution studies with synchrotron radiation, hydrogen atoms of proteins are clearly observed. In addition, outer-shell electrons of proteins can be analyzed by multipolar refinement procedures. (Fig. 2)
Miki says, "I am very thankful that, throughout my career, I have experienced the enormous progress of protein crystallography as a research method and witnessed the new technology development that might lead to a breakthrough in this field.”
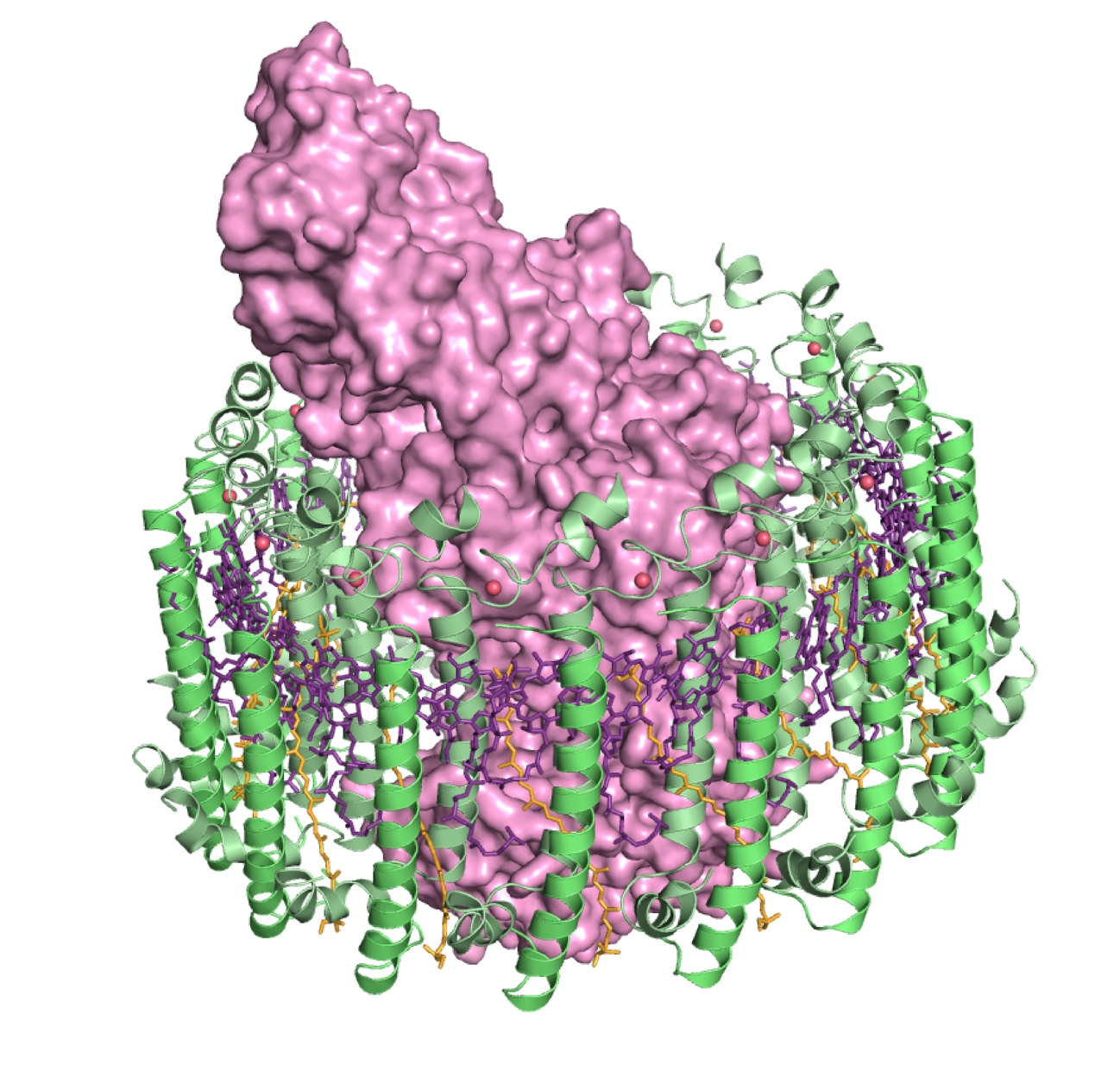
Figure 1
Prof. Miki’s research group found the light-harvesting core antenna (LH1) and the reaction center (RC) of photosynthetic bacteria form a supramolecular complex (LH1–RC) to use sunlight energy in a highly efficient manner. (S. Niwa et al., Nature 2014) It took 30 years to elucidate this crystal structure after the photosynthetic RC structure was determined. (J. Deisenhofer et al., Nature 1985)
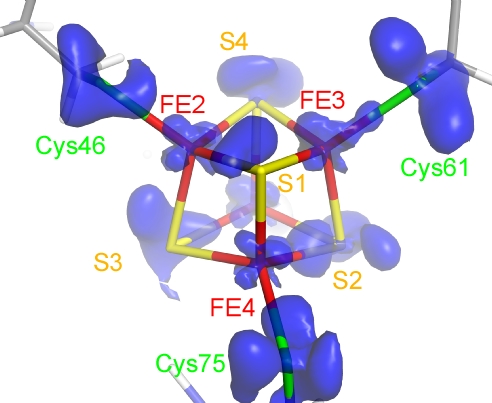
Figure 2
Outer-shell electron density of the Fe4S4 cluster in the crystal structure at 0.48 Å resolution of high-potential iron-sulfur protein (HiPIP) of photosynthetic bacteria. (Y. Hirano et al., Nature 2016)



